Multimorbidity Patterns
Hidden Markov Models
“Recently, several advanced machine-learning techniques such as non-hierarchical and hierarchical clustering techniques have been used to explore multimorbidity patterns.”
Hidden Markov Models (HMM) were developed based on the Bayesian Information Criterion. The Bayesian Information Criterion is an algorithm of inference that is used to select the best model from a set of possible models. It is a powerful technique for analyzing temporal data that can capture dynamic changes in longitudinal patterns over time. Since HMMs can account for complex longitudinal data, they are well-suited to investigate the dynamics of multimorbidity over time.
The Study
In this study, HMMs were used to investigate 3,363 older adults from the Swedish National study on Aging and Care in Kungsholmen (SNAC-K) and the evolution of their multimorbidity patterns over the course of 12 years. The aim of this research was to explore the evolution of these patterns across decades of life in older adults and to examine how they transition across different chronic diseases when further chronic diseases arise. In this cohort of study participants, the average age was 76.1 years old, 66.6% were female and 87.2% had multimorbidity at baseline.
The researchers divided the participants into three groups based on age: sexagenarians (between 60 and 66 years), septuagenarians (between 72 and 78 years) and octogenarians (81 years and over). Data used in the HMMs included age, gender, education level, self-reported chronic diseases and medications, results from a Mini Mental State Examination (MMSE), and walking speed. Data were collected from participants at baseline and at six and 12 years (three separate time points).
“At each follow-up wave, SNAC-K participants undergo an approximately five-hour-long comprehensive clinical and functional assessment carried out by trained physicians, nurses, and neuropsychologists.”
Results
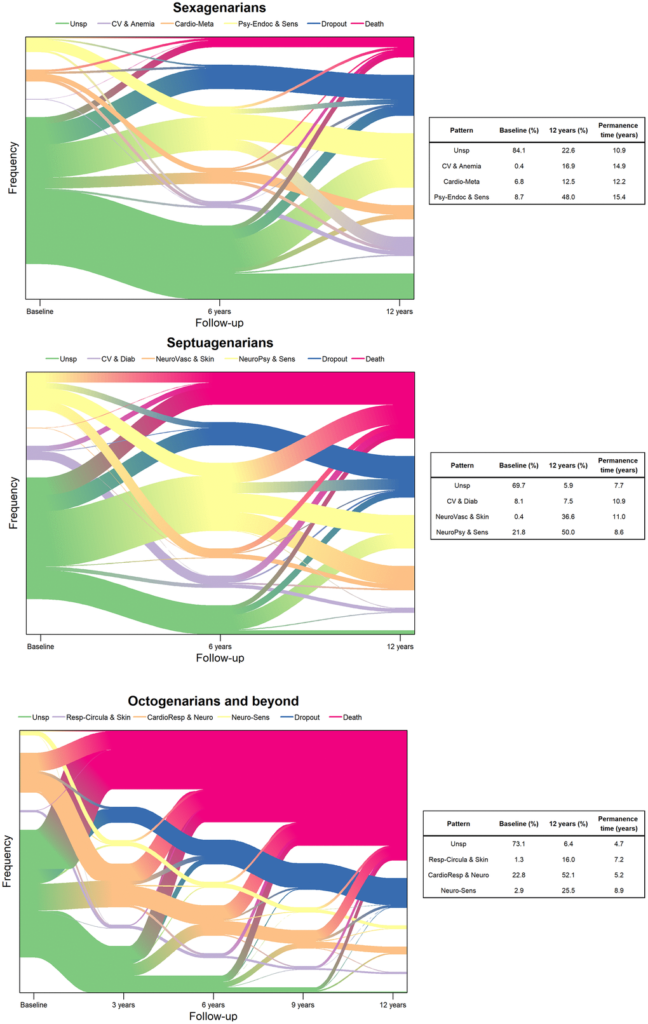
The team identified four longitudinal multimorbidity patterns in each decade. The Unspecific pattern consists of participants with no specific pattern of multimorbidity. In all decades, participants showed the shortest permanence time in the Unspecific pattern. The researchers also included categories for participants who dropped out of the study or passed away.
Next, the top 10 diseases were selected out of each age group at each follow-up wave to identify the most common multimorbidity patterns. Among the sexagenarians, the multimorbidity patterns were clustered into cardiovascular and anemia, cardio-metabolic, and psychiatric-endocrine and sensorial. Among the septuagenarians, the multimorbidity patterns were clustered into cardiovascular and diabetes, neuro-vascular and skin-sensorial, and neuro-psychiatric and sensorial. Among the octogenarians, the multimorbidity patterns were clustered into respiratory-circulatory and skin, cardio-respiratory and neurological, and neuro-sensorial. The data showed that participants commonly shifted from one pattern to another. (See Figure 1.)
“In this study we identified and characterized longitudinal multimorbidity patterns among older adults from a Swedish urban population, and estimated the time they spent in each pattern as well as the probability of transitioning across different patterns throughout a 12-year follow-up period.”
Conclusions
“Our statistical approach enabled us to model the evolution and transitions of multimorbidity over time, and the results of this could be applied in the interests of healthier aging. Moreover, the age-stratified analyses allowed us to identify which disease combinations and transitions were more prevalent in each decade.”
The findings of this study suggest that multimorbidity patterns change with age and highlight the importance of understanding the dynamic nature of multimorbidity over time. Through the use of HMMs, this research was able to detect changes in the prevalence and transition of multimorbidity patterns across different decades of life. These findings can help healthcare providers and researchers better understand the complex nature of multimorbidity and develop more effective interventions for older adults. Furthermore, this research provides evidence that the use of HMMs to study longitudinal data is a useful tool for further research into multimorbidity. Additional studies with more data is needed to gain a better understanding of the interplay between multimorbidity and aging.
“Our study provides evidence that multimorbidity is dynamic and heterogeneous in old age. With increasing age, older adults experience decreasing clinical stability and progressively shorter permanence time within one same multimorbidity pattern. Moreover, a significant proportion ranging between 5.9%-22.6% belongs to an Unspecific pattern with a low burden of diseases and a promising preventive potential. Adding new variables related to drug use, environmental and genetic factors, and/or frailty to the longitudinal analysis of multimorbidity patterns may allow optimizing the epidemiological understanding and applicability of these models for patient-tailored prevention and management strategies.”
Click here to read the full research paper published by Aging.
AGING (AGING-US) VIDEOS: YouTube | LabTube | Aging-US.com
—
Aging is an open-access journal that publishes research papers bi-monthly in all fields of aging research. These papers are available at no cost to readers on Aging-us.com. Open-access journals have the power to benefit humanity from the inside out by rapidly disseminating information that may be freely shared with researchers, colleagues, family, and friends around the world.
For media inquiries, please contact [email protected].